biometrics
What is biometrics?
Biometrics is the measurement and statistical analysis of people's unique physical and behavioral characteristics. The technology is mainly used for identification and access control or for identifying individuals who are under surveillance. The basic premise of biometric authentication is that every person can be accurately identified by intrinsic physical or behavioral traits. The term biometrics is derived from the Greek words bio, meaning life, and metric, meaning to measure.
How do biometrics work?
Authentication by biometric verification is becoming increasingly common in corporate and public security systems, consumer electronics and point-of-sale applications. In addition to security, the driving force behind biometric verification has been convenience, as there are no passwords to remember or security tokens to carry. Some biometric methods, such as measuring a person's gait, can operate with no direct contact with the person being authenticated.
Components of biometric devices include the following:
- a reader or scanning device to record the biometric factor being authenticated;
- software to convert the scanned biometric data into a standardized digital format and to compare match points of the observed data with stored data; and
- a database to securely store biometric data for comparison.
Biometric data may be held in a centralized database, although modern biometric implementations often depend instead on gathering biometric data locally and then cryptographically hashing it so that authentication or identification can be accomplished without direct access to the biometric data itself.
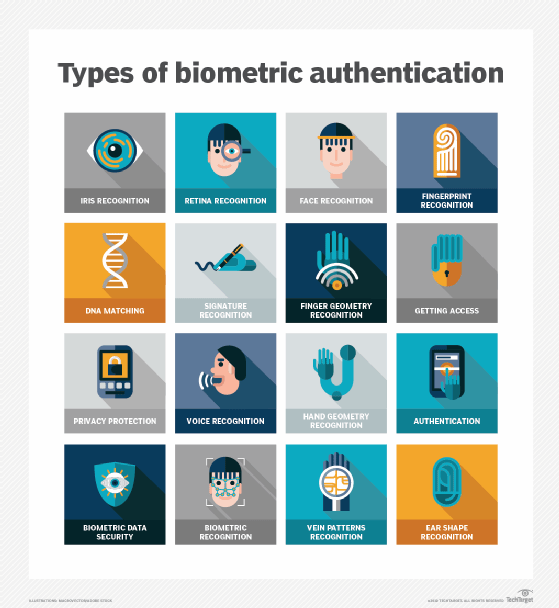
Types of biometrics
The two main types of biometric identifiers are either physiological characteristics or behavioral characteristics.
Physiological identifiers relate to the composition of the user being authenticated and include the following:
- facial recognition
- fingerprints
- finger geometry (the size and position of fingers)
- iris recognition
- vein recognition
- retina scanning
- voice recognition
- DNA (deoxyribonucleic acid) matching
- digital signatures
Behavioral identifiers include the unique ways in which individuals act, including recognition of typing patterns, mouse and finger movements, website and social media engagement patterns, walking gait and other gestures. Some of these behavioral identifiers can be used to provide continuous authentication instead of a single one-off authentication check. While it remains a newer method with lower reliability ratings, it has the potential to grow alongside other improvements in biometric technology.
Biometric data can be used to access information on a device like a smartphone, but there are also other ways biometrics can be used. For example, biometric information can be held on a smart card, where a recognition system will read an individual's biometric information, while comparing that against the biometric information on the smart card.
Advantages and disadvantages of biometrics
The use of biometrics has plenty of advantages and disadvantages regarding its use, security and other related functions. Biometrics are beneficial for the following reasons:
- hard to fake or steal, unlike passwords;
- easy and convenient to use;
- generally, the same over the course of a user's life;
- nontransferable; and
- efficient because templates take up less storage.
Disadvantages, however, include the following:
- It is costly to get a biometric system up and running.
- If the system fails to capture all of the biometric data, it can lead to failure in identifying a user.
- Databases holding biometric data can still be hacked.
- Errors such as false rejects and false accepts can still happen.
- If a user gets injured, then a biometric authentication system may not work -- for example, if a user burns their hand, then a fingerprint scanner may not be able to identify them.
Examples of biometrics in use
Aside from biometrics being in many smartphones in use today, biometrics are used in many different fields. As an example, biometrics are used in the following fields and organizations:
- Law enforcement. It is used in systems for criminal IDs, such as fingerprint or palm print authentication systems.
- United States Department of Homeland Security. It is used in Border Patrol branches for numerous detection, vetting and credentialing processes -- for example, with systems for electronic passports, which store fingerprint data, or in facial recognition systems.
- Healthcare. It is used in systems such as national identity cards for ID and health insurance programs, which may use fingerprints for identification.
- Airport security. This field sometimes uses biometrics such as iris recognition.
However, not all organizations and programs will opt in to using biometrics. As an example, some justice systems will not use biometrics so they can avoid any possible error that may occur.
What are security and privacy issues of biometrics?
Biometric identifiers depend on the uniqueness of the factor being considered. For example, fingerprints are generally considered to be highly unique to each person. Fingerprint recognition, especially as implemented in Apple's Touch ID for previous iPhones, was the first widely used mass-market application of a biometric authentication factor.
Other biometric factors include retina, iris recognition, vein and voice scans. However, they have not been adopted widely so far, in some part, because there is less confidence in the uniqueness of the identifiers or because the factors are easier to spoof and use for malicious reasons, like identity theft.
Stability of the biometric factor can also be important to acceptance of the factor. Fingerprints do not change over a lifetime, while facial appearance can change drastically with age, illness or other factors.
The most significant privacy issue of using biometrics is that physical attributes, like fingerprints and retinal blood vessel patterns, are generally static and cannot be modified. This is distinct from nonbiometric factors, like passwords (something one knows) and tokens (something one has), which can be replaced if they are breached or otherwise compromised. A demonstration of this difficulty was the over 20 million individuals whose fingerprints were compromised in the 2014 U.S. Office of Personnel Management data breach.
The increasing ubiquity of high-quality cameras, microphones and fingerprint readers in many of today's mobile devices means biometrics will continue to become a more common method for authenticating users, particularly as Fast ID Online has specified new standards for authentication with biometrics that support two-factor authentication with biometric factors.
While the quality of biometric readers continues to improve, they can still produce false negatives, when an authorized user is not recognized or authenticated, and false positives, when an unauthorized user is recognized and authenticated.
Are biometrics secure?
While high-quality cameras and other sensors help enable the use of biometrics, they can also enable attackers. Because people do not shield their faces, ears, hands, voice or gait, attacks are possible simply by capturing biometric data from people without their consent or knowledge.
An early attack on fingerprint biometric authentication was called the gummy bear hack, and it dates back to 2002 when Japanese researchers, using a gelatin-based confection, showed that an attacker could lift a latent fingerprint from a glossy surface. The capacitance of gelatin is similar to that of a human finger, so fingerprint scanners designed to detect capacitance would be fooled by the gelatin transfer.
Determined attackers can also defeat other biometric factors. In 2015, Jan Krissler, also known as Starbug, a Chaos Computer Club biometric researcher, demonstrated a method for extracting enough data from a high-resolution photograph to defeat iris scanning authentication. In 2017, Krissler reported defeating the iris scanner authentication scheme used by the Samsung Galaxy S8 smartphone. Krissler had previously recreated a user's thumbprint from a high-resolution image to demonstrate that Apple's Touch ID fingerprinting authentication scheme was also vulnerable.
After Apple released iPhone X, it took researchers just two weeks to bypass Apple's Face ID facial recognition using a 3D-printed mask; Face ID can also be defeated by individuals related to the authenticated user, including children or siblings.






